About the CRISPR/Cas9 system
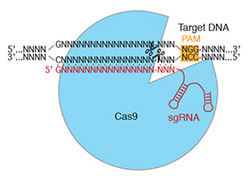
Cas9 is an endonuclease that functions in bacterial adaptive immunity. In nature, Cas9 binds to two small RNAs: a trans-activating RNA (tracrRNA) that activates its enzymatic activity, and a CRISPR RNA (crRNA) that determines substrate specificity. In a seminal paper published in 2012, Jinek, Chylinski and colleagues showed that the crRNA recognizes substrates by base pairing with the target DNA. In addition, they found that the tracrRNA and crRNA could be combined into a chimeric single guide RNA (sgRNA) that supported target DNA cleavage with high efficiency and specificity. There are only two requirements for Cas9 to cleave a substrate: base pairing between the 5' end of the sgRNA and the target DNA, and the presence of an NGG motif (the PAM) immediately downstream of the target sequence. Thus, by changing the targeting sequence at the 5' end of the sgRNA, Cas9 can be programmed to target a wide variety of sequences.
This amazing discovery set off a wave of papers reporting the use of Cas9 for genome editing in a wide variety of organisms including human cells, mice, flies, yeast, bacteria... and our favorite model organism, the nematode C. elegans.
This amazing discovery set off a wave of papers reporting the use of Cas9 for genome editing in a wide variety of organisms including human cells, mice, flies, yeast, bacteria... and our favorite model organism, the nematode C. elegans.
About Cas9-triggered homologous recombination
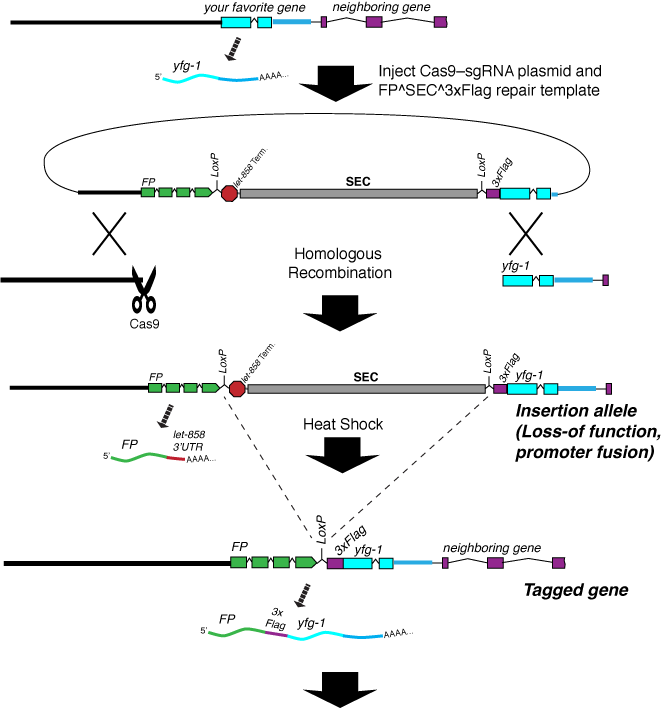
Cas9-triggered homologous recombination is a fast and inexpensive way to make essentially any desired change to the C. elegans genome. The principle is simple: we use Cas9 to make a DNA double-strand break at a defined site in the genome. At the same time, we introduce a homologous repair template containing the genome modification we want to make. The double-strand break is repaired by homologous recombination with the modified template we supply. In this way, we can generate insertions, deletions, point mutants, in-frame GFP fusions, or any other modification we can imagine.
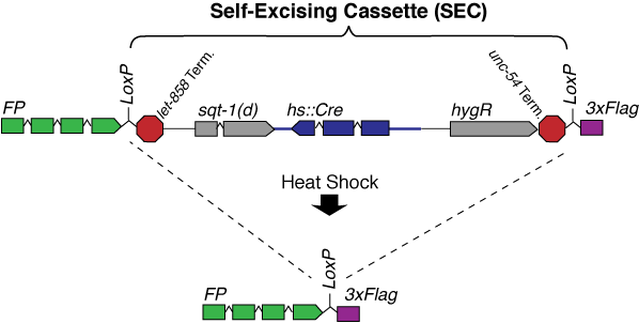
To streamline the editing process, we recently published a new protocol that makes use of a Self-Excising Cassette (SEC) for drug selection. SEC contains three parts: 1) A Hygromycin resistance gene (hygR) for positive selection in any genetic background; 2) A dominant phenotypic marker (sqt-1(d)) that confers a Roller phenotype; and 3) An inducible Cre recombinase (hs::Cre) that excises itself from the genome, along with hygR and sqt-1(d), upon heat shock. SEC is flanked by LoxP sites and placed within an intron of a fluorescent protein (FP)::3xFlag tag, so that after marker removal, the residual LoxP site is with an intron of the FP tag.
To streamline the editing process, we recently published a new protocol that makes use of a Self-Excising Cassette (SEC) for drug selection. SEC contains three parts: 1) A Hygromycin resistance gene (hygR) for positive selection in any genetic background; 2) A dominant phenotypic marker (sqt-1(d)) that confers a Roller phenotype; and 3) An inducible Cre recombinase (hs::Cre) that excises itself from the genome, along with hygR and sqt-1(d), upon heat shock. SEC is flanked by LoxP sites and placed within an intron of a fluorescent protein (FP)::3xFlag tag, so that after marker removal, the residual LoxP site is with an intron of the FP tag.
With an appropriate Cas9 plasmid and homology arms, an FP^SEC^3xFlag module can be inserted anywhere in the C. elegans genome: at the N-terminus, C-terminus, or in place of any gene of interest. If FP^SEC^3xFlag is inserted at the N-terminus of a gene, our workflow produces a loss-of-function allele, a promoter fusion and a protein fusion in a single injection step. For more details, see our paper in Genetics and the Protocols page).
Our approach builds on previous genome modification strategies for C. elegans, especially the work of Christian Frøkjaer-Jensen and the Bessereau lab. We've also benefited from discussions with the Seydoux lab and with Jordan Ward. We are grateful to these colleagues for their contributions to the field and to our work in particular.